The BMI lab bridges research in Machine Learning and Sequence Analysis methodology research and its application to biomedical problems. We collaborate with biologists and clinicians to develop real-world solutions.
We work on research questions and foundational challenges in storing, analysing, and searching extensive heterogeneous and temporal data, especially in the biomedical domain. Our lab members address technical and non-technical research questions in collaboration with biologists and clinicians. At the research group’s core is an active knowledge exchange in both directions between the methods and the application-driven researchers.
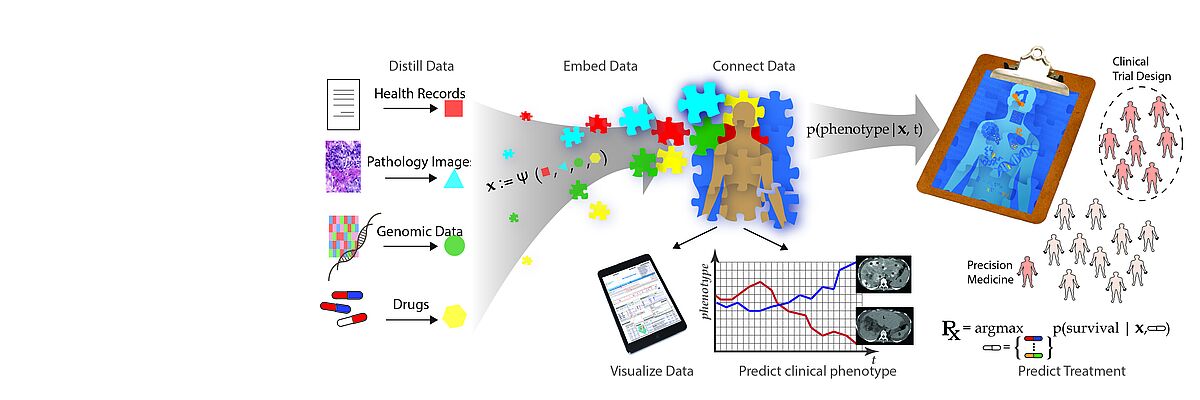
The emergence of data-driven medicine leverages data and algorithms to shape how we diagnose and treat patients. Machine Learning approaches allow us to capitalise on the vast amount of data produced in clinical settings to generate novel biomedical insights and build more precise predictive models of disease outcomes and treatment efficacy.
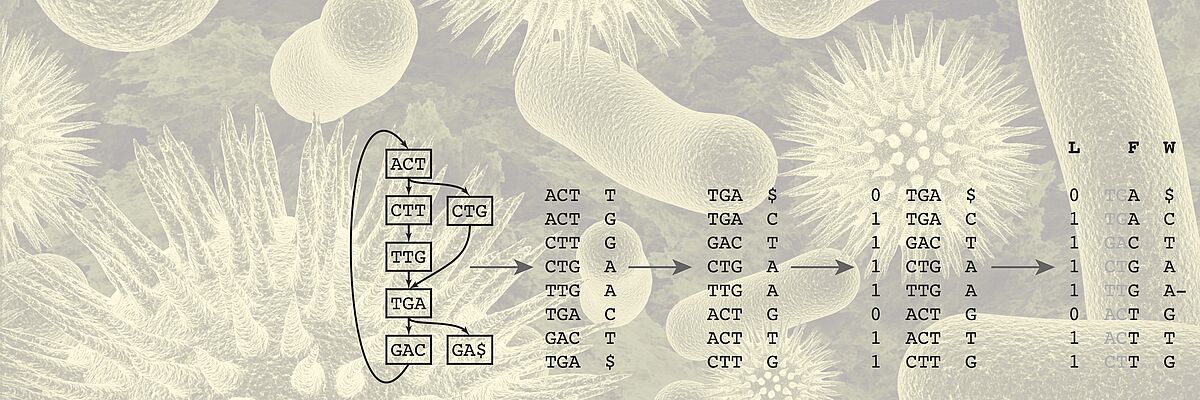
We work towards this transformation mainly but not exclusively in two key areas. One key application area is the analysis of heterogeneous data of cancer patients. For Genomics, we develop algorithms for storing, compressing, and searching extensive genomics datasets. Another key area is the development of time series models of patient health states and early warning systems for intensive care units.
Publications
Abstract Spatial transcriptomics technology remains resource-intensive and unlikely to be routinely adopted for patient care soon. This hinders the development of novel precision medicine solutions and, more importantly, limits the translation of research findings to patient treatment. Here, we present DeepSpot, a deep-set neural network that leverages recent foundation models in pathology and spatial multi-level tissue context to effectively predict spatial transcriptomics from H&E images. DeepSpot substantially improved gene correlations across multiple datasets from patients with metastatic melanoma, kidney, lung, or colon cancers as compared to previous state-of-the-art. Using DeepSpot, we generated 1 792 TCGA spatial transcriptomics samples (37 million spots) of the melanoma and renal cell cancer cohorts. We anticipate this to be a valuable resource for biological discovery and a benchmark for evaluating spatial transcriptomics models. We hope that DeepSpot and this dataset will stimulate further advancements in computational spatial transcriptomics analysis.
Authors Kalin Nonchev, Sebastian Dawo, Karina Selina, Holger Moch, Sonali Andani, Tumor Profiler Consortium, Viktor Hendrik Koelzer, and Gunnar Rätsch
Submitted MedRxiv
Abstract Notable progress has been made in generalist medical large language models across various healthcare areas. However, large-scale modeling of in-hospital time series data - such as vital signs, lab results, and treatments in critical care - remains underexplored. Existing datasets are relatively small, but combining them can enhance patient diversity and improve model robustness. To effectively utilize these combined datasets for large-scale modeling, it is essential to address the distribution shifts caused by varying treatment policies, necessitating the harmonization of treatment variables across the different datasets. This work aims to establish a foundation for training large-scale multi-variate time series models on critical care data and to provide a benchmark for machine learning models in transfer learning across hospitals to study and address distribution shift challenges. We introduce a harmonized dataset for sequence modeling and transfer learning research, representing the first large-scale collection to include core treatment variables. Future plans involve expanding this dataset to support further advancements in transfer learning and the development of scalable, generalizable models for critical healthcare applications.
Authors Manuel Burger, Fedor Sergeev, Malte Londschien, Daphné Chopard, Hugo Yèche, Eike Gerdes, Polina Leshetkina, Alexander Morgenroth, Zeynep Babür, Jasmina Bogojeska, Martin Faltys, Rita Kuznetsova, Gunnar Rätsch
Submitted Best Paper @ NeurIPS AIM-FM Workshop 2024
Abstract Fatigue is one of the most prevalent symptoms of chronic diseases, such as Multiple Sclero- sis, Alzheimer’s, and Parkinson’s. Recently researchers have explored unobtrusive and continuous ways of fatigue monitoring using mobile and wearable devices. However, data quality and limited labeled data availability in the wearable health domain pose significant challenges to progress in the field. In this work, we perform a systematic evaluation of self-supervised learning (SSL) tasks for fatigue recognition using wearable sensor data. To establish our benchmark, we use Homekit2020, which is a large-scale dataset collected using Fitbit devices in everyday life settings. Our results show that the majority of the SSL tasks outperform fully supervised baselines for fatigue recognition, even in limited labeled data scenarios. In particular, the domain fea- tures and multi-task learning achieve 0.7371 and 0.7323 AUROC, which are higher than the other SSL tasks and supervised learning baselines. In most of the pre-training tasks, the performance is higher when using at least one data augmentation that reflects the potentially low quality of wearable data (e.g., missing data). Our findings open up promising opportunities for continuous assessment of fatigue in real settings and can be used to guide the design and development of health monitoring systems.
Authors Tamás Visy, Rita Kuznetsova, Christian Holz, Shkurta Gashi
Submitted CHIL 2024 (PMLR)
Abstract Spatial transcriptomics enables in-depth molecular characterization of samples on a morphology and RNA level while preserving spatial location. Integrating the resulting multi-modal data is an unsolved problem, and developing new solutions in precision medicine depends on improved methodologies. Here, we introduce AESTETIK, a convolutional deep learning model that jointly integrates spatial, transcriptomics, and morphology information to learn accurate spot representations. AESTETIK yielded substantially improved cluster assignments on widely adopted technology platforms (e.g., 10x Genomics™, NanoString™) across multiple datasets. We achieved performance enhancement on structured tissues (e.g., brain) with a 21% increase in median ARI over previous state-of-the-art methods. Notably, AESTETIK also demonstrated superior performance on cancer tissues with heterogeneous cell populations, showing a two-fold increase in breast cancer, 79% in melanoma, and 21% in liver cancer. We expect that these advances will enable a multi-modal understanding of key biological processes.
Authors Kalin Nonchev, Sonali Andani, Joanna Ficek-Pascual, Marta Nowak, Bettina Sobottka, Tumor Profiler Consortium, Viktor Hendrik Koelzer, and Gunnar Rätsch
Submitted MedRxiv
Abstract This study advances Early Event Prediction (EEP) in healthcare through Dynamic Survival Analysis (DSA), offering a novel approach by integrating risk localization into alarm policies to enhance clinical event metrics. By adapting and evaluating DSA models against traditional EEP benchmarks, our research demonstrates their ability to match EEP models on a time-step level and significantly improve event-level metrics through a new alarm prioritization scheme (up to 11% AuPRC difference). This approach represents a significant step forward in predictive healthcare, providing a more nuanced and actionable framework for early event prediction and management.
Authors Hugo Yèche, Manuel Burger, Dinara Veshchezerova, Gunnar Rätsch
Submitted CHIL 2024